Mycobacterium phage Lucky13
Know something about this phage that we don't? Modify its data.
| Detailed Information for Phage Lucky13 | |
| Discovery Information | |
| Isolation Host | Mycobacterium smegmatis mc²155 |
| Found By | Robert S. Malloy |
| Year Found | 2010 |
| Location Found | Fort Payne, AL USA |
| Finding Institution | Jacksonville State University |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | Yes |
| Isolation Temperature | Not entered |
| GPS Coordinates | 33.872777 N, 85.939166 W Map |
| Discovery Notes | The sample was collected at Little River Canyon in Fort Payne, AL. The sample came from the root of the Never Wet Plant. |
| Sequencing Information | |
| Sequencing Complete? | No |
| Genome length (bp) | Unknown |
| Character of genome ends | Unknown |
| Fasta file available? | No |
| Characterization | |
| Cluster | Unclustered |
| Subcluster | -- |
| Annotating Institution | Unknown or unassigned |
| Annotation Status | Not sequenced |
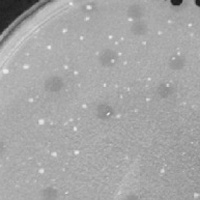
| Plaque Notes | Plating of the original sample resulted in two distinct morphologies. My partner, Stacey Grimes, and I separated them. Lucky 13 formed clear plaques with an average diameter of 2 mm. |
| Has been Phamerated? | No |
| Publication Info | |
| Uploaded to GenBank? | No |
| GenBank Accession | None yet |
| Refseq Number | None yet |
| Archiving Info | |
| Archiving status | Archived |
| SEA Designator | 2010JASTlucky13RSM |
| Pitt Freezer Box# | 14 |
| Pitt Freezer Box Grid# | 75 |
| Available Files | |
| Plaque Picture | Download |
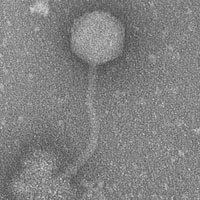
| EM Picture | Download |