| Detailed Information for Phage Prager | |
| Discovery Information | |
| Isolation Host | Mycobacterium smegmatis mc²155 |
| Found By | Jackelyn Prager |
| Year Found | 2023 |
| Location Found | Pittsburgh, PA USA |
| Finding Institution | University of Pittsburgh |
| Program | Phage Hunters Integrating Research and Education |
| From enriched soil sample? | Yes |
| Isolation Temperature | 37°C |
| GPS Coordinates | 40.444638 N, 79.952252 W Map |
| Discovery Notes | This phage was isolated from a soil sample collected by Jackie Prager on 11/8/2023. Phage isolation and purification were performed by Deborah Jacobs-Sera. Jackie, a producer at CNN, was here to document phage hunting for a CNN International piece. |
| Naming Notes | Jackie named it after herself! |
| Sequencing Information | |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Feb 7, 2024 |
| Sequencing Facility | Pittsburgh Bacteriophage Institute |
| Shotgun Sequencing Method | Illumina Sequencing |
| Approximate Shotgun Coverage | 10485 |
| Genome length (bp) | 64462 |
| Character of genome ends | Circularly Permuted |
| GC Content | 59.7% |
| Fasta file available? | Yes: Download fasta file |
| Characterization | |
| Cluster | D |
| Subcluster | D1 |
| Cluster Life Cycle | Lytic |
| Other Cluster Members | |
| Annotating Institution | University of Pittsburgh |
| Annotation Status | Being Annotated (Expected completion by 5/1/2024) |
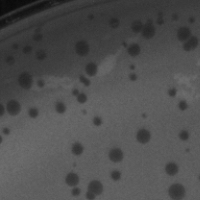
| Plaque Notes | The plaues are tiny, ~2mm, and realtively clear. |
| Morphotype | Siphoviridae |
| Has been Phamerated? | Yes |
| Gene List |
Click to ViewPrager_Draft_1 Prager_Draft_2 Prager_Draft_3 Prager_Draft_4 Prager_Draft_5 Prager_Draft_6 Prager_Draft_7 Prager_Draft_8 Prager_Draft_9 Prager_Draft_10 Prager_Draft_11 Prager_Draft_12 Prager_Draft_13 Prager_Draft_14 Prager_Draft_15 Prager_Draft_16 Prager_Draft_17 Prager_Draft_18 Prager_Draft_19 Prager_Draft_20 Prager_Draft_21 Prager_Draft_22 Prager_Draft_23 Prager_Draft_24 Prager_Draft_25 Prager_Draft_26 Prager_Draft_27 Prager_Draft_28 Prager_Draft_29 Prager_Draft_30 Prager_Draft_31 Prager_Draft_32 Prager_Draft_33 Prager_Draft_34 Prager_Draft_35 Prager_Draft_36 Prager_Draft_37 Prager_Draft_38 Prager_Draft_39 Prager_Draft_40 Prager_Draft_41 Prager_Draft_42 Prager_Draft_43 Prager_Draft_44 Prager_Draft_45 Prager_Draft_46 Prager_Draft_47 Prager_Draft_48 Prager_Draft_49 Prager_Draft_50 Prager_Draft_51 Prager_Draft_52 Prager_Draft_53 Prager_Draft_54 Prager_Draft_55 Prager_Draft_56 Prager_Draft_57 Prager_Draft_58 Prager_Draft_59 Prager_Draft_60 Prager_Draft_61 Prager_Draft_62 Prager_Draft_63 Prager_Draft_64 Prager_Draft_65 Prager_Draft_66 Prager_Draft_67 Prager_Draft_68 Prager_Draft_69 Prager_Draft_70 Prager_Draft_71 Prager_Draft_72 Prager_Draft_73 Prager_Draft_74 Prager_Draft_75 Prager_Draft_76 Prager_Draft_77 Prager_Draft_78 Prager_Draft_79 Prager_Draft_80 Prager_Draft_81 Prager_Draft_82 Prager_Draft_83 Prager_Draft_84 Prager_Draft_85 Prager_Draft_86 Prager_Draft_87 |
| Publication Info | |
| Uploaded to GenBank? | No |
| GenBank Accession | None yet |
| Refseq Number | None yet |
| Archiving Info | |
| Archiving status | Not in Pitt Archives |
| Available Files | |
| Plaque Picture | Download |