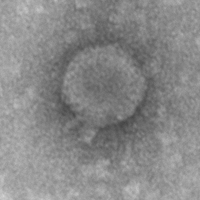
| Detailed Information for Phage Stella |
| Discovery Information |
| Isolation Host | Streptomyces viridochromogenes DSM40736 |
| Found By | Kylie Mink and Guanlan Dong |
| Year Found | 2015 |
| Location Found | Clayton, MO USA |
| Finding Institution | Washington University in St. Louis |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | No |
| Isolation Temperature | 29°C |
| GPS Coordinates | 38.643893 N, 90.315244 W Map |
| Discovery Notes | The soil sample was taken from a patch of ivy underneath a small tree outside of a dorm building window. The soil was fairly dry and was found in an urban setting. |
| Naming Notes | We named the phage Stella due to the star-shaped plaques the phage formed. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Jul 21, 2022 |
| Sequencing Facility | The McDonnell Genome Institute at Washington University |
| Shotgun Sequencing Method | Illumina |
| Sequencer Used | Illumina MiSeq |
| Read Type | Single-end reads |
| Read Length | 150 bp |
| Approximate Shotgun Coverage | 226 |
| Genome length (bp) | 46511 |
| Character of genome ends | Direct Terminal Repeat |
| Direct Terminal Repeat Length | 263 bp |
| GC Content | 60.0% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | BF |
| Subcluster | -- |
| Cluster Life Cycle | Lytic |
| Other Cluster Members |
|
| Annotating Institution | Washington University in St. Louis |
| Annotation Status | In GenBank |
| Plaque Notes | After five rounds of purification, we obtained a plate with plaques that were all the same medium size. The plaques were star-shaped with a round, clear center and cloudy, rough edges. |
| Morphotype | Podoviridae |
| Number of Genes | 69 |
| Number of tRNAs | 21 |
| Number of tmRNAs | 0 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Minimal DNA Master File | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | OP751154 |
| Refseq Number | None yet |
| Archiving Info |
| Archiving status |
Not in Pitt Archives |
| SEA Designator |
Stella |
| SEA Lysate Titer |
2.1 x 10^8 pfu/mL |
| Date of SEA Lysate Titering |
Oct 13, 2015 |
| Available Files |
| GenBank File for Phamerator | Download |
