| Detailed Information for Phage DS6A |
| Discovery Information |
| Isolation Host | Mycobacterium tuberculosis H37Rv |
| Found By | W. B. Redmond |
| Year Found | 1960 |
| Location Found | Unavailable |
| Finding Institution | University of Pittsburgh |
| Program | Phage Hunters Integrating Research and Education |
| From enriched soil sample? | Unknown |
| Isolation Temperature | Not entered |
| GPS Coordinates | 40.444800 N, 79.952700 W Map |
| Discovery Notes | Original Citation: W.B. Redmond and J.C. Cater "A bacteriophage specific for Mycobacterium tuberculosis, varieties hominis and bovis" Amer. Rev. Resp. Dis. 82:781-786 (1960).
Sample obtained from William R. Jacobs lab for sequencing.
Originally isolated from stockyard soil. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Sep 24, 2010 |
| Genome length (bp) | 60588 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 12 bases |
| Overhang Sequence | CGAGGCCGACAT |
| GC Content | 68.4% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | Singleton |
| Subcluster | -- |
| Cluster Life Cycle | Unknown |
| Other Cluster Members |
|
| Annotating Institution | Unknown or unassigned |
| Annotation Status | In GenBank |
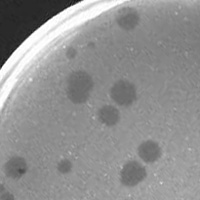
| Plaque Notes | Large plaques (see pic.). Infects all and only mycobacteria of the TB complex (M. tuberculosis, M. bovis, BCG, M. africanum, M. microti, etc.). The Hatfull lab has tested the lysate on M. smegmatis and confirmed that it does not infect it. |
| Number of Genes | 97 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | JN698994 |
| Refseq Number | NC_023744 |
| Published in Paper | Pope, Bowman, Russell, et al 2015, eLIFE: Whole genome comparison of a large collection of mycobacteriophages reveals a continuum of phage genetic diversity |
| Archiving Info |
| Archiving status |
Archived |
| Pitt Freezer Box# |
1 |
| Pitt Freezer Box Grid# |
F12 |
| Available Files |
| Plaque Picture | Download |
| Virtual Digest Picture | Download |
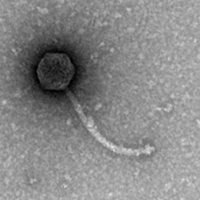
| EM Picture | Download |
| Final DNAMaster File | Download |