| Discovery Information |
| Isolation Host | Gordonia rubripertincta NRRL B-16540 |
| Found By | Jimmy Vo |
| Year Found | 2021 |
| Location Found | Jeffersonville, IN United States |
| Finding Institution | Indiana University-Southeast |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | Yes |
| Isolation Temperature | 26°C |
| GPS Coordinates | 38.314609 N, 85.693118 W Map |
| Discovery Notes | The soil sample was found in front of my house next to a bush and where I put my trash. I picked a spot with little grass and needed a shovel due to how tough and dry the soil was. Once in a bag, it was mushy, moist, and soft. |
| Naming Notes | The bacteriophage was named Survivor because the spot test used after Enriched Isolation was contaminated. There was also a large spot of contamination next to the sample. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Jan 8, 2022 |
| Sequencing Facility | Pittsburgh Bacteriophage Institute |
| Shotgun Sequencing Method | Illumina |
| Sequencer Used | Illumina MiSeq |
| Read Type | Single-end reads |
| Read Length | 150 bp |
| Approximate Shotgun Coverage | 735 |
| Genome length (bp) | 45436 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 10 bases |
| Overhang Sequence | CGGTAGGCAT |
| GC Content | 61.8% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | CT |
| Subcluster | -- |
| Cluster Life Cycle | Lytic |
| Other Cluster Members |
|
| Annotating Institution | Indiana University-Southeast |
| Annotation Status | In GenBank |
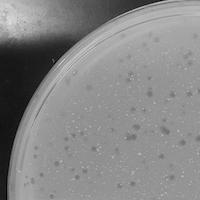
| Plaque Notes | When there is a little number of plaques on the plaque assay, they have spread far apart and are hard to see. The morphology is cloudy with no clear or consistent shape like a small dot. The diameter is half a millimeter. However, when highly concentrated the plaques are clumped together in one area and are clear. The plaques are dark with a perfectly round shape. |
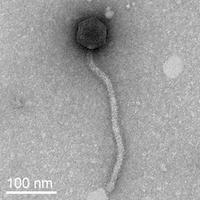
| Morphotype | Siphoviridae |
| Number of Genes | 69 |
| Number of tRNAs | 0 |
| Number of tmRNAs | 0 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Annotation in DNA Master Format | Download |
| Submitted Minimal DNA Master File | Download |
| Author List | Download |
| Annotation Cover Sheet | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | ON970576 |
| Refseq Number | None yet |
| Archiving Info |
| Archiving status |
Archived |
| Pitt Freezer Box# |
139 |
| Pitt Freezer Box Grid# |
F3 |
| Available Files |
| GenBank File for Phamerator | Download |