| Detailed Information for Phage Werner |
| Discovery Information |
| Isolation Host | Streptomyces lividans JI 1326 |
| Found By | Neal Huang and Ethan Cordes |
| Year Found | 2019 |
| Location Found | St. Louis, MO United States |
| Finding Institution | Washington University in St. Louis |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | No |
| Isolation Temperature | 28°C |
| GPS Coordinates | 38.6352 N, 90.2687 W Map |
| Discovery Notes | We named the Werner phage after Werner Heisenberg, specifically because of his famous uncertainty principle. While working with the Werner phage we were always unable to determine whether our solutions actually had any phage particles in them. Without this confirmation, we were always uncertain whether or not our experiments would succeed. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Jun 24, 2020 |
| Sequencing Facility | The McDonnell Genome Institute at Washington University |
| Shotgun Sequencing Method | Illumina Sequencing |
| Approximate Shotgun Coverage | 100 |
| Genome length (bp) | 51566 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 11 bases |
| Overhang Sequence | CGGCCAGTGAT |
| GC Content | 65.7% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | BD |
| Subcluster | BD1 |
| Cluster Life Cycle | Temperate |
| Other Cluster Members |
|
| Annotating Institution | Washington University in St. Louis |
| Annotation Status | In GenBank |
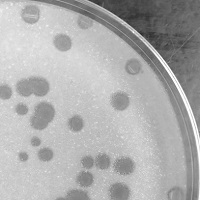
| Plaque Notes | The plaques have a bullseye pattern where the edges are more turbid, and the centers are clearer. The plaques are also ringed in a red pigment and are approximately 3.96mm in diameter. The plaques displayed in the included image were grown on a 100mm plate and were produced using a phage solution diluted to a concentration of 10^5. |
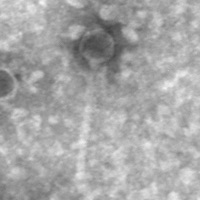
| Morphotype | Siphoviridae |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Minimal DNA Master File | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | MW435856 |
| Refseq Number | None yet |
| Archiving Info |
| Archiving status |
Not in Pitt Archives |
| Available Files |
| Plaque Picture | Download |
| Restriction Digest Picture | Download |
| EM Picture | Download |
| GenBank File for Phamerator | Download |