
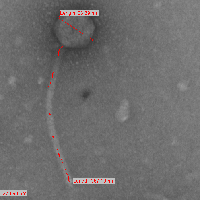
| Detailed Information for Phage Stark |
| Discovery Information |
| Isolation Host | Mycobacterium smegmatis mc²155 |
| Found By | John Park |
| Year Found | 2012 |
| Location Found | Baltimore, MD USA |
| Finding Institution | Johns Hopkins University |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | Yes |
| Isolation Temperature | Not entered |
| GPS Coordinates | 39.2986 N, 76.5946 W Map |
| Discovery Notes | The soil used was from a tomato plant patch. It was relatively wet. It also had various insects living in the patch. |
| Naming Notes | The name stark comes from one of my favorite tv series, the Game of Thrones. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Feb 6, 2013 |
| Sequencing Facility | Virginia Commonwealth University Nucleic Acids Research Facilities |
| Shotgun Sequencing Method | 454 |
| Sequencer Used | 454 Genome Sequencer FLX |
| Approximate Shotgun Coverage | 140 |
| Sanger Finishing Reactions | 12 |
| Genome length (bp) | 74731 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 9 bases |
| Overhang Sequence | CGCTTGTCA |
| GC Content | 63.0% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | E |
| Subcluster | -- |
| Cluster Life Cycle | Temperate |
| Other Cluster Members |
|
| Annotating Institution | University of Pittsburgh |
| Annotation Status | In GenBank |
| Plaque Notes | Relatively small, Fuzzy on the outer areas of the plaque. about 1.5 mm diameter. |
| Morphotype | Siphoviridae |
| Number of Genes | 141 |
| Number of tRNAs | 2 |
| Number of tmRNAs | 0 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Minimal DNA Master File | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | MN586019 |
| Refseq Number | None yet |
| Published in Paper | Pope, Bowman, Russell, et al 2015, eLIFE: Whole genome comparison of a large collection of mycobacteriophages reveals a continuum of phage genetic diversity |
| Archiving Info |
| Archiving status |
Archived |
| Pitt Freezer Box# |
13 |
| Pitt Freezer Box Grid# |
C9 |
| Available Files |
| Plaque Picture | Download |
| EM Picture | Download |
| Final DNAMaster File | Download |
| GenBank File for Phamerator | Download |