| Detailed Information for Phage Murica |
| Discovery Information |
| Isolation Host | Mycobacterium smegmatis mc²155 |
| Found By | Team Miley Virus |
| Year Found | 2013 |
| Location Found | Los Angeles, CA USA |
| Finding Institution | University of California, Los Angeles |
| Program | Unavailable |
| From enriched soil sample? | Yes |
| Isolation Temperature | 35°C |
| GPS Coordinates | 34.07 N, 118.451 W Map |
| Discovery Notes | Phage isolation performed in Fall 2013 and genome annotation performed in Spring 2014 by Team Miley Virus: Kola Awe, Dami Oshinuga, and Apryl Stafford. Annotation quality checked by Lei Yeh, Edric Yoon, and Iger Ostreni. TA: Samantha Capati. Instructor: Dr. Jordan Moberg Parker. DNA preparation performed by Krisanavane Reddi (UCLA). Assembly performed by William Villella (UCLA).
Moist soil collected next to a mossy tree, foot traffic and regular sprinklers on October 3, 2013. |
| Naming Notes | Slang for America. |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Apr 11, 2014 |
| Sequencing Facility | The UCLA Genotyping and Sequencing Core |
| Shotgun Sequencing Method | 454 |
| Sequencer Used | 454 Genome Sequencer FLX |
| Approximate Shotgun Coverage | 81 |
| Genome length (bp) | 77053 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 9 bases |
| Overhang Sequence | CGCTTGTCA |
| GC Content | 63.0% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | E |
| Subcluster | -- |
| Cluster Life Cycle | Temperate |
| Other Cluster Members |
|
| Annotating Institution | Unknown or unassigned |
| Annotation Status | In GenBank |
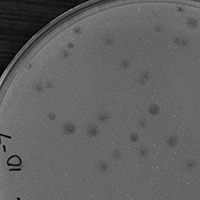
| Plaque Notes | Murica plaque assays contain both completely clear plaques in addition to plaques with clear centers and turbid edges. Average size (diameter in mm): 3.5mm. Range (in mm): 3mm-5mm |
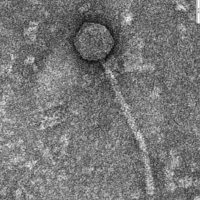
| Morphotype | Siphoviridae |
| Number of Genes | 147 |
| Number of tRNAs | 2 |
| Number of tmRNAs | 0 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Minimal DNA Master File | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | MF919525 |
| Refseq Number | None yet |
| Archiving Info |
| Archiving status |
Archived |
| Pitt Freezer Box# |
23 |
| Pitt Freezer Box Grid# |
C6 |
| Available Files |
| Plaque Picture | Download |
| Restriction Digest Picture | Download |
| EM Picture | Download |
| GenBank File for Phamerator | Download |