
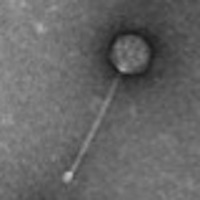
| Detailed Information for Phage Nibb |
| Discovery Information |
| Isolation Host | Mycobacterium smegmatis mc²155 |
| Found By | Alex Neitz and Vanessa Villela |
| Year Found | 2014 |
| Location Found | Spokane, WA USA |
| Finding Institution | Gonzaga University |
| Program | Science Education Alliance-Phage Hunters Advancing Genomics and Evolutionary Science |
| From enriched soil sample? | Yes |
| Isolation Temperature | 25°C |
| GPS Coordinates | 47.667233 N, 117.402333 W Map |
| Discovery Notes | Collected after light rain, found 2 in deep in soil, date found 9/3/14 at 10:43 am, 11.11 degrees celius outside temperature |
| Sequencing Information |
| Sequencing Complete? | Yes |
| Date Sequencing Completed | Aug 23, 2016 |
| Sequencing Facility | Pittsburgh Bacteriophage Institute |
| Shotgun Sequencing Method | Illumina |
| Sequencer Used | Illumina MiSeq |
| Read Type | Single-end reads |
| Read Length | 150 bp |
| Approximate Shotgun Coverage | 1486 |
| Genome length (bp) | 62293 |
| Character of genome ends | 3' Sticky Overhang |
| Overhang Length | 11 bases |
| Overhang Sequence | CTCGTAGGCAT |
| GC Content | 67.6% |
| Fasta file available? | Yes: Download fasta file |
| Characterization |
| Cluster | K |
| Subcluster | K1 |
| Cluster Life Cycle | Temperate |
| Other Cluster Members |
|
| Annotating Institution | Gonzaga University |
| Annotation Status | In GenBank |
| Plaque Notes | Small, circular, turbid on edges, and concentration dependent |
| Morphotype | Siphoviridae |
| Number of Genes | 102 |
| Number of tRNAs | 1 |
| Number of tmRNAs | 0 |
| Has been Phamerated? | Yes |
| Gene List |
|
| Submitted Minimal DNA Master File | Download |
| Publication Info |
| Uploaded to GenBank? | Yes |
| GenBank Accession | MK460246 |
| Refseq Number | None yet |
| Archiving Info |
| Archiving status |
Archived |
| SEA Lysate Titer |
2.145e10 |
| Date of SEA Lysate Titering |
Oct 16, 2014 |
| Pitt Freezer Box# |
16 |
| Pitt Freezer Box Grid# |
H10 |
| Available Files |
| Plaque Picture | Download |
| Restriction Digest Picture | Download |
| EM Picture | Download |
| GenBank File for Phamerator | Download |